Statistics
For more comprehensive information on ecological statistics please see Magurran [6] and Colwell [3].
A. Alpha Diversity
Alpha diversity indices measure the sequence/OTU diversity of individual Explicet libraries. When comparing alpha diversity metrics between libraries with different sequence counts, rarefaction is essential. Rarefaction is a process that randomly resamples all libraries to the sampling depth of the smallest library (called the rarefaction point), thus allowing the alpha diversity metrics to be compared directly (i.e., by equalizing sequencing effort across libraries). The curves generated by plotting the alpha diversity metrics versus sample size are called collectors curves and are indicative of how well-sampled libraries are. Species observed (Sobs), which simply counts the number of OTUs in a library, provides a good intuitive example of how to interpret collectors curves. If a Sobs graph has a steep upwards slope at the terminal end of the curve, it indicates that additional taxa are likely to be found with a small amount of additional sampling effort. If the Sobs curve flattens out, a sufficient number of sequences have likely been taken as no more taxa will be found with additional sequencing.
Tools → Analyze → Alpha Diversity
- A new pop-up window will appear
1. Settings
a) Libraries
The user can select whether to calculate diversity for all the libraries, a subset of libraries, or groups of libraries by clicking on the various options at the top and right of the pop-up window. For descriptions of these options, see Display.
b) Bootstrap
Explicet performs rarefaction based analysis of alpha diversity measure through bootstrapping, which allows estimation of both an index and its variability through repeated sub-sampling of a dataset. Clicking on the Bootstrap button causes Explicet to start the random resampling process. At the rarefaction point, bootstrap resampling is conducted at the size of the smallest library, thus allowing comparison of biodiversity indices between libraries under the condition of equal sampling effort.
- Default: 25. The dataset will be resampled 25 times.
c) Cutoff Size
Varies by dataset. The default will select all libraries ≥ the 3rd sigma of the average library size value. Some very small libraries may be excluded, which will be displayed directly to the right of the Cutoff Size box, labeled by # Libs Exc..
d) # Steps
In generating a collector’s curve, the step size controls the incremental increase in number of sequences between resampling events, in other words the number of data points in each curve. The number of sequences in the smallest included library is divided by # Steps to create the step size. This step size will then be applied to all the larger libraries for calculations and rarefaction. The number of sequences per step is displayed to the right of the # Steps box.
- Default: 0 steps.
By default, Explicet calculates a Single statistic at Rarefaction point only. To remove this default, deselect the checkbox below the # Steps box.
When # Steps is set at the default (0 steps) and Single statistic at Rarefaction point only is unchecked, # Steps will automatically change to 10.
Note: As the # Steps increases, Bootstrap will take longer to run. To create smoother curves, increase # Steps, but keep in mind that the length of time to run Bootstrap will increase exponentially.
e) Use Min Lib Size
Using the minimum library size is necessary for comparison of multiple curves/libraries. Most studies will have this option checked; Use Min Lib Size should be checked to reduce bias. For example, if you have one library with 500 sequences and another library with 10,000 sequences, the value at rarefaction is where the smallest library size falls; this way, the libraries are compared at the same level of sampling effort.
You may choose to un-check Use Min Lib Size to analyze individual libraries irrespective of the size of other libraries. Without Use Min Lib Size checked, sampling starts at each library’s total Counts divided by 10 (when # Steps: 10). Note that if this option is not selected, rarefaction will not apply due to sampling variation/sampling bias between libraries. Setting # Steps to 1 and un-checking Use Min Lib Size will result in calculation of diversity indices for the total number of sequences in each library. In other words, rarefaction will not be performed.
f) User Modifications
- The user may modify any of the default settings to be more or less stringent.
2. Calculate
After selecting all the appropriate settings, click Bootstrap on upper right side of window. This will generally take a few seconds to minutes depending on computer processing speed. You will be unable to use other Explicet functions while the program is calculating alpha diversity.
3. Tests
a) Sobs
- The number of taxa/OTUs observed in a library.
b) Singletons
- The number of taxa/OTUs observed exactly once in a library.
c) Doubletons
- The number of taxa/OTUS observed exactly twice in a library.
d) ACE
- Abundance based Coverage Estimator. The number of predicted taxa/OTUs based on observed singletons and rare taxa.
e) ACEVar
- Coefficient of variation of the Abundance based Coverage Estimator.
f) Chao1
- The number of species predicted based on observed singletons and doubletons.
g) Chao195ciL
- Chao1 lower 95% confidence interval based on the bootstrap.
h) Chao195ciU
- Chao1 upper 95% confidence interval based on the bootstrap.
i) Goods
- A measure of how well the amount of sequencing done for a library represents the biodiversity in the library. Low coverage means that the library is under-sampled and may require additional sequencing. “Good’s Coverage” is measured on a 0-100% scale, with 100% indicating that all expected OTUs have been observed.
j)ShannonH
- Shannon diversity index, H (log base 2). This measure of biodiversity incorporates both OTU richness and OTU evenness into a single value. Greater OTU count and more uniform distribution of OTUs increase H.
k) ShannonE
- Shannon evenness index, H/Hmax (%). Measures the uniformity of a distribution of OTUs on a 0-100% scale. A perfectly uniform distribution will have E = 100%.
l) Simpson
- Simpson’s index, D: Another measure of complexity that measures the probability that two randomly selected individuals belong to the same OTU.
m) SimpsonD
- Simpson’s diversity index: 1 – D.
n) SimpsonE
- Simpson’s evenness index.
o) SimpsonR
- The reciprocal of Simpson’s index: 1/D.
4. Plot
After calculating the alpha diversity, click Plot on lower right side of window. A pop-up will appear with all the diversity measures and library names. Select the desired parameters, then click OK to plot the rarefaction curves for the selected measure and libraries. Toggle the display of individual curves on and off by clicking on the legend. Click Remove Deselected Curves to remove the curves from the figure. See Figures for details on how to modify titles, axis labels, and colors.
B. Beta Diversity
- Beta diversity indices measure the similarity (or dissimilarity) in microbiome composition between pairs of libraries. A number of such indices, which differ in their weighting of abundant and rarer OTUs, have been proposed:
1. Indices
a) Morisita-Horn
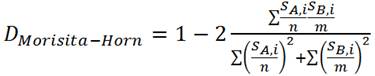
SA,i = the number of individuals from community A in the ith OTU
SB,i = the number of individuals from community B in the ith OTU
- n = the number of individuals in community A
- m = the number of individuals in community B
(Formula from Mothur website: http://www.mothur.org/wiki/Morisitahorn)
- Morisita-Horn is a beta diversity measure on a 0-1 scale, indicating relative similarity between OTUs contained within two libraries; 1 = identical overlap in OTUs between libraries, 0 = no shared OTUs between two libraries.
Tools → Analyze → Beta Diversity → Morisita-Horn
- A new window will appear that contains a comparison matrix with the calculated Morisita-Horn values for each pair of libraries or metadata groupings. A value of 1.000 indicates two libraries are identical, which is why the diagonal of the matrix (libraries compared to themselves) has values of 1.000.
b) Bray-Curtis
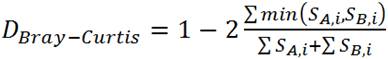
SA,i = the number of individuals in the ith OTU of community A
SB,i = the number of individuals in the ith OTU of community B
(Formula from Mothur website: http://www.mothur.org/wiki/Braycurtis)
- Bray-Curtis is a beta diversity measure on a 0-1 scale, indicating relative dissimilarity between OTUs contained within two libraries; 0 = identical overlap in OTUs between libraries, 1 = no shared OTUs between two libraries.
Tools → Analyze → Beta Diversity → Bray-Curtis
- A new window will appear that contains a comparison matrix with the calculated Bray-Curtis values for each pair of libraries or metadata groupings. A value of 0.000 indicates two groups are identical, which is why the diagonal of the matrix (libraries compared to themselves) has values of 0.000.
c) ThetaYC
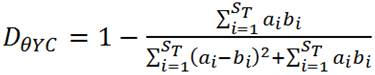
ST = the total number of OTUs in communities A and B
ai = the relative abundance of OTU i in community A
bi = the relative abundance of OTU i in community B
(Formula from Mothur website: http://www.mothur.org/wiki/Thetayc)
- ThetaYC is a beta diversity measure on a 0-1 scale, indicating relative dissimilarity between OTUs contained within two libraries; 0 = identical overlap in OTUs between libraries, 1 = no shared OTUs between two libraries.
Tools → Analyze → Beta Diversity → ThetaYC
- A new window will appear that contains a comparison matrix with the calculated ThetaYC values for each pair of libraries or metadata groupings. A value of 0.000 indicates two groups are identical, which is why the diagonal of the matrix (libraries compared to themselves) has values of 0.000.
2. Display Options
The matrix display, and subsequent plots, can be modified by rearranging libraries to group by higher values (column total of each library) in either the upper left (Descending by Value) or lower right (Ascending by Value) corner. The default order is Alphabetical by Library Name, which can be useful if libraries within a metadata group are defined by similar names.
3. Plot
A heatmap of the matrix is generated by clicking Plot on the lower right side of window. See the section on heatmaps (Figures) for information on modifying the plot attributes.
C. Two-Part Test
This tool provides a statistical test of whether individual OTUs differ significantly in abundance and/or prevalence between two groups of data. The Two-Part test is the sum of two test statistics, one comparing the proportion of non-zero counts and one comparing the medians of the non-zero counts. The statistical approaches incorporated in Explicet are useful for identification of taxa that differentiate two groups. The Two-Part statistic makes use of a common practice in statistics by adding the Chi2 statistics from two independent tests. This was developed to address the issue of inverse relationship between proportion of samples that contained a taxon (i.e., taxon prevalence) and the median relative abundance of that taxon. For more information on the Two-Part test, please see: Wagner et al [11].
The input requires that the user select two mutually exclusive categories of libraries (e.g., healthy and diseased) using filters. To create filters see Select Data. The developers recommend that at least five libraries be in each category so that the test is statistically valid.
Tools → Analyze → Two-Part
1. Settings
a) Select Filters
The number of libraries included in each filter will be displayed immediately to the right of the filter selection. If no filters have been created, click Setup Filters on upper right side of window and proceed as described in Select Data.
b) Select P-Threshold
- Default: 0, which will show the P-value for every OTU.
c) OTU Display
Select the desired OTU name display options, see Display for details.
2. Output
- The output table displays the following variables with values for each of the taxa: m1, p1, med1, m2, p2, med2, chi2 (chi**2), PValue and -Log(PV)…
a) m1 and m2
- The number of samples with non-zero sequence counts for each category.
b) p1 and p2
- The % of non-zero counts for each corresponding group: p1 and p2 are the prevalences of an OTU in the two categories.
c) med1 and med2
- The median of the non-zero sequence counts (or median of the “m” samples).
d) chi**2
- The chi-squared test statistic for the two-part test.
e) PValue
- The corresponding parametric p-value from the two-part test.
f) -Log(PV)
- The negative of the log(base 10) of the p-value. This transformation is used to more easily visualize those taxa that are statistically significant in a Manhattan plot.
3. Plot
Once Two-Part statistics have been calculated, the data can be visualized as a Manhattan plot (scatterplot with lines) by clicking Plot on lower right side of window. The x-axis is the OTU number from the Two-Part spreadsheet; the y-axis is the -Log(PV). The plots have three horizontal marker lines. The bottom line indicates results approaching significance at a p-value of 0.1. The middle line denotes significance at a p-value of 0.05, and the top line denotes significance at a p-value of 0.01.
See Figures for details on how to modify titles, axis labels, and colors.
D. Two-Proportions Test
- The Two-Proportions test assesses the differences in proportions of individual OTUs between two populations. This comparative tool performs a continuity-adjusted chi-square test to evaluate the difference in the detection rates across the two groups for each taxon/OTU.
The input requires the user to apply two mutually exclusive filters to the libraries tested. To create filters, see Select Data.
Tools → Analyze → Two-Proportions
1. Settings
a) Select Filters
The number of libraries included in each filter will be displayed immediately to the right of the filter selection. If no filters have been created, click Setup Filters on upper right side of window and proceed as described in Select Data.
b) Select P-Threshold
- Default: 0, which will show the P-value for every OTU.
c) OTU Display
Select the desired OTU name display options, see Display for details.
2. Output
- The output table displays the following variables with values for each of the taxa: m1, p1, m2, p2, Z, PValue and -Log(PV)…
a) m1 and m2
- The number of samples with non-zero sequence counts for each corresponding group.
b) p1 and p2
- The % of non-zero counts for each corresponding group.
c) Z
- The two-proportions test statistic.
d) PValue
- The corresponding parametric p-value from the two-proportions test.
e) -Log(PV)
- The negative of the log(base 10) of the p-value.
3. Plot
Once Two-Proportion statistics have been calculated, the data can be visualized as a Manhattan plot (scatterplot with lines) by clicking Plot on lower right side of window. The x-axis is the OTU number from the Two-Proportion spreadsheet; the y-axis is the -Log(PV). The plots have three horizontal marker lines. The bottom line indicates results approaching significance at a p-value of 0.1. The middle line denotes significance at a p-value of 0.05, and the top line denotes significance at a p-value of 0.01.
See Figures for details on how to modify titles, axis labels, and colors.
E. Wilcoxon Test
- The Wilcoxon test is a nonparametric statistical calculation used to compare the median OTU abundances of two categories. A non-parametric Wilcoxon test with a continuity correction is employed because OTU abundances are not necessarily normally distributed. This version of the Wilcoxon test compares the median relative abundance from all libraries in a pair of treatment groups. This differs from the Wilcoxon test used in the Two-Part test, which includes only libraries with median relative abundances greater than 0.0.
The input requires the user to apply two mutually exclusive filters to the libraries tested. To create filters, see Select Data.
Tools → Analyze → Wilcoxon
1. Settings
a) Select Filters
The number of libraries included in each filter will be displayed immediately to the right of the filter selection. If no filters have been created, click Setup Filters on upper right side of window and proceed as described in Select Data.
b) Select P-Threshold
- Default: 0, which will show the P-value for every OTU.
c) OTU Display
Select the desired OTU name display options, see Display for details.
2. Output
- The output table displays the following variables with values for each of the taxa: m1, n1, p1, med1, m2, n2, p2, med2, W, PValue and -Log(PV)…
a) m1 and m2
- The number of samples with non-zero sequence counts for each corresponding category.
b) n1 and n2
- The total number of samples in each category.
c) p1 and p2
- The % of non-zero counts for each corresponding category.
d) med1 and med2
- The median of the non-zero sequence counts (or median of the “m” samples).
e) W
- The Wilcoxon value.
f) PValue
- The parametric p-value from the two-part test.
g) -Log(PV)
- The negative of the log(base 10) of the p-value. This transformation provides a convenient means of plotting p-values, such as in a Manhattan plot.
3. Plot
Once Wilcoxon statistics have been calculated, the data can be visualized as a Manhattan plot (scatterplot with lines) by clicking Plot on lower right side of window. The x-axis is the OTU number from the Wilcoxon spreadsheet; the y-axis is the -Log(PV). The plots have three horizontal marker lines. The bottom line indicates results approaching significance at a p-value of 0.1. The middle line denotes significance at a p-value of 0.05, and the top line denotes significance at a p-value of 0.01.
See Figures for details on how to modify titles, axis labels, and colors.